3@brief `main()` shows how to use [cross_correlate()](#locate_signal.cross_correlate)
4and [plot_correlation_map()](#locate_signal.plot_correlation_map).
6\anchor example_one_pair_corr_map
7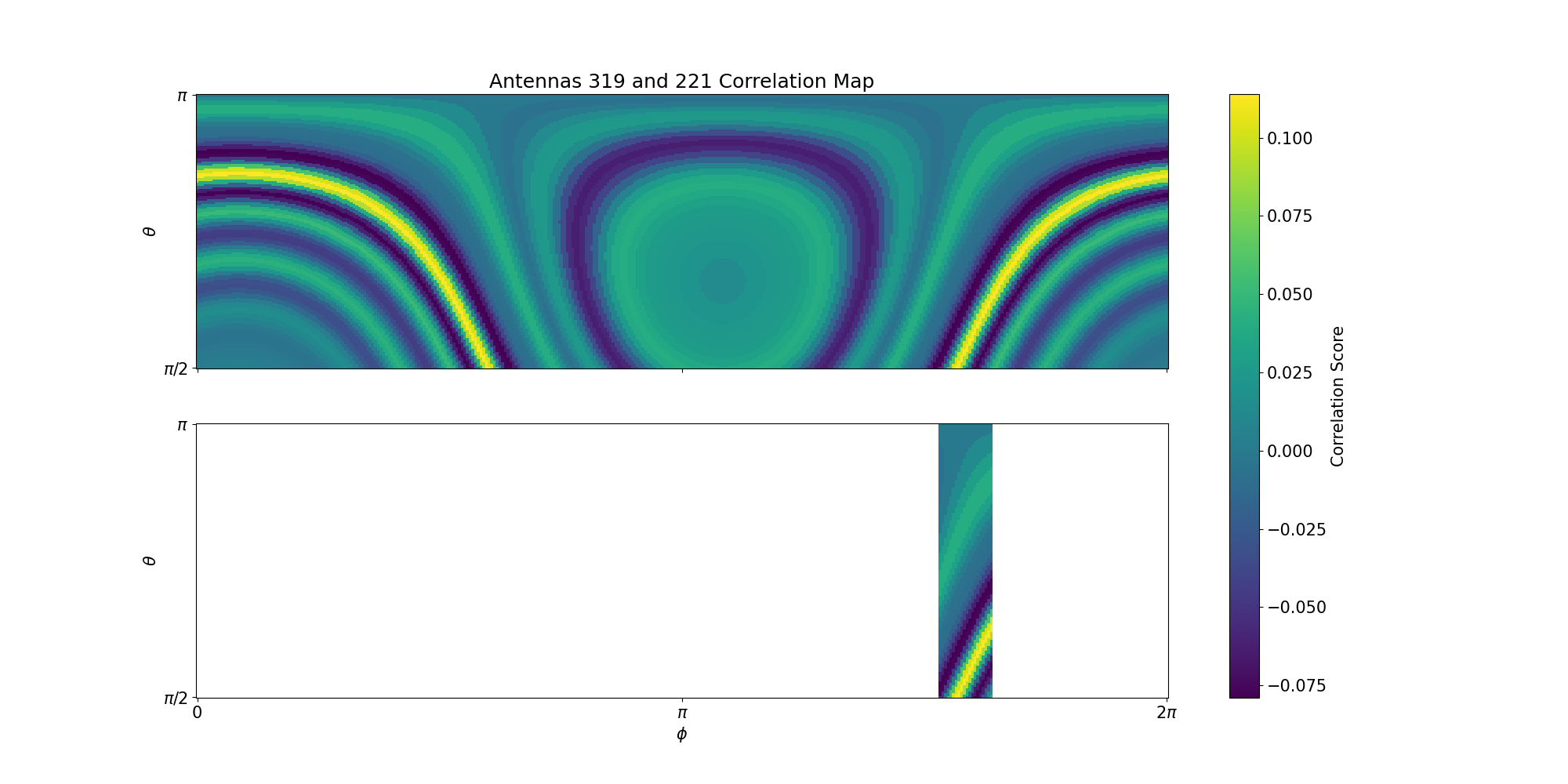
9Summing over all antenna pairs, we have
11\anchor example_corr_map
12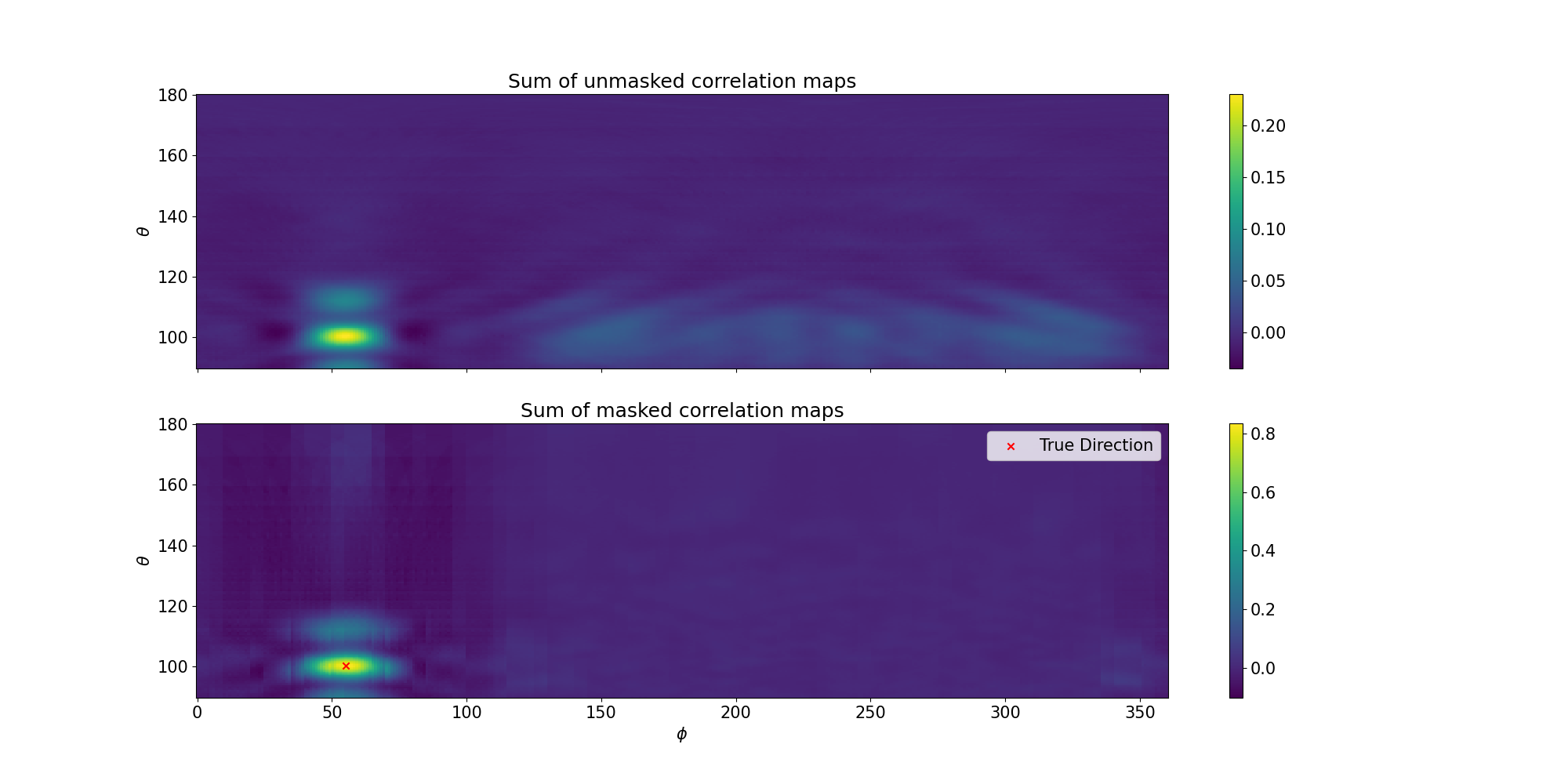
14We can see that by masking the individual correlation maps, the total correlation map is less
23from time_delays
import _make_grid
24from pathlib
import Path
25from initialise
import load_pueoEvent_Dataset
35def supply_azimuthal_angle_masks(skymaps: pl.DataFrame) -> pl.DataFrame:
36 r"""!Supplies azimuth masks to any sky map
37 (eg [time delay maps](#time_delays.make_time_delay_skymap) or
38 [correlation maps](#cross_correlate))
40 @param[in] skymaps columns `A1_PhiSector` and `A2_PhiSector` are **required**
41 @retval phi_masks see the schema of the example output below
44 * Input: required columns are the \f$\phi\f$-sectors of the antenna pairs,
45 `A1_PhiSector` and `A2_PhiSector`.
46 * Output schema: one column (`masks`) will be attached to the input dataframe,
50$ masks <array[bool, (1, 360)]>
55 * \f$\phi\f$-sector 1 is centered around 0 degrees azimuth, \f$\phi\f$-sector 2 around 15 degrees,
56 and so on and so forth; the last \f$\phi\f$ sector (24) is centered around 345 degrees.
57 * Based on the antenna's [field of view](#antenna_attributes.ASSUMED_PHI_SECTOR_APERTURE_WIDTH),
58 the values outside a certain range are dropped.
59 * In the figure below, we show what this would look like for antennas from the first three and
60 the last \f$\phi\f$-sectors (white means masked).
62 
64 * The chosen masking behavior for now is that **only the data within the overlapping unmasked
65 range would be kept**.
66 * For instance, suppose an antenna from \f$\phi\f$-sector 1 is paired with an antenna from
67 \f$\phi\f$-sector 2, then:
69 
72 * Thus, if the antenna pair's
73 [fields of view](#antenna_attributes.ASSUMED_PHI_SECTOR_APERTURE_WIDTH)
74 do not overlap, then everything will be masked.
76 from antenna_attributes
import NUM_PHI_SECTORS, ASSUMED_PHI_SECTOR_APERTURE_WIDTH
77 phi, _ = _make_grid(sparse=
True)
84 {
"PhiSector": np.arange(1, 25)}, schema={
"PhiSector": pl.UInt8}
88 ((pl.col(
"PhiSector") - 1) * (360 / NUM_PHI_SECTORS)).alias(
"aperture center [deg]")
92 pl.col(
"aperture center [deg]"),
94 (pl.col(
"aperture center [deg]") - ASSUMED_PHI_SECTOR_APERTURE_WIDTH / 2)
95 ).alias(
"left bound [deg]"),
97 (pl.col(
"aperture center [deg]") + ASSUMED_PHI_SECTOR_APERTURE_WIDTH / 2)
98 ).alias(
"right bound [deg]"),
105 aperture_bounds.select(
107 pl.col(
"left bound [deg]").alias(
"A1 left"),
108 pl.col(
"right bound [deg]").alias(
"A1 right"),
109 ), left_on=
"A1_PhiSector", right_on=
"PhiSector"
111 aperture_bounds.select(
113 pl.col(
"left bound [deg]").alias(
"A2 left"),
114 pl.col(
"right bound [deg]").alias(
"A2 right"),
115 ), left_on=
"A2_PhiSector", right_on=
"PhiSector"
117 pl.struct(pl.col(
"A1 left"), pl.col(
"A1 right"))
121 ((phi - left) % 360) < ((right - left) % 360)
126 pl.struct(pl.col(
"A2 left"), pl.col(
"A2 right"))
130 ((phi - left) % 360) < ((right - left) % 360)
138 "A1 masks",
"A2 masks",
"A1 left",
"A2 left",
"A1 right",
"A2 right"
140 pl.struct(
"A1 masks",
"A2 masks").map_batches(
142 ~(m1 & m2)
for m1, m2
in s.to_numpy()
150def combine_time_delay_maps_and_waveforms(masks_and_skymaps: pl.DataFrame,
151 waveforms: pl.DataFrame) -> pl.DataFrame:
152 r"""!Prepares a big table that has all the column needed by #cross_correlate.
154 @param[in] masks_and_skymaps The output of #supply_azimuthal_angle_masks
155 @param[in] waveforms The output of #waveform_plots.load_waveforms
156 @retval big_frame See sample output schema below.
159 * The following columns are required in `waveforms`:
161 -# `waveforms (volts)`
162 -# `step size (nanoseconds)`
164 * The following columns are required in `masks_and_skymaps`:
167 -# `time delays [sec]`
174$ A1_waveforms (volts) <array[f64, 3072]>
175$ A2_waveforms (volts) <array[f64, 3072]>
176$ time delays [samples] <array[i64, (180, 360)]>
177$ masks <array[bool, (1, 360)]>
180 * `time delays [samples]` refers to the [time delay skymaps](#time_delays.make_time_delay_skymap),
181 with the units converted from seconds to "samples"
182 * That is, the units are in "steps" (`step size (nanoseconds)`)
183 * The values in these time delay skymaps are therefore integers, serving as indices.
184 * These indices are then used later in #cross_correlate when creating the
185 correlation skymaps based on the time delay maps (via "fancy-indexing").
186 * Qualitatively, the time delay maps have not changed. For example:
187 \image html time_delay_map_in_seconds.svg Time Delay Map in Seconds width=40%
188 \image html time_delay_map_in_samples.svg Time Delay Map in Samples width=40%
191 signal_length = len(waveforms[
"waveforms (volts)"][0])
192 step_size = waveforms[
"step size (nanoseconds)"][0] * 1e-9
196 .join(waveforms, left_on=
"A1_AntNum", right_on=
"AntNum")
197 .join(waveforms, left_on=[
"A2_AntNum",
"Pol"], right_on=[
"AntNum",
"Pol"])
200 pl.col(
"time delays [sec]") / step_size + signal_length - 1
203 .map_batches(
lambda s: np.rint(s.to_numpy()).astype(int))
204 .alias(
"time delays [samples]")
207 pl.col(
"waveforms (volts)").alias(
"A1_waveforms (volts)"),
208 pl.col(
"waveforms (volts)_right").alias(
"A2_waveforms (volts)")
211 r"^A[12]_AntNum$",
"Pol",
r"^A[12]_waveforms \(volts\)$",
"time delays [samples]",
"masks"
218def cross_correlate(big_frame: pl.DataFrame) -> pl.DataFrame:
219 r"""!Compute the zero-centered normalized cross correlation (ZNCC) and makes correlation skymaps
221 @param[in] big_frame The output of #combine_time_delay_maps_and_waveforms()
222 @retval correlation_maps See the schema of the example output below
224 * Parameters: columns `correlation` and `correlation maps` will be added to the input,
225 so the output schema looks like
230$ A1_waveforms (volts) <array[f64, 3072]>
231$ A2_waveforms (volts) <array[f64, 3072]>
232$ time delays [samples] <array[i64, (180, 360)]>
233$ masks <array[bool, (1, 360)]>
234$ correlation <array[f64, 6143]>
235$ correlation maps <array[f64, (180, 360)]>
237 * `correlation maps` contain correlation skymaps.
238 These are matrices with the same dimensions as the time delay maps of `time delays [samples]`,
239 as the former are made based on the latter via "fancy-indexing"
241 * Each matrix element of a correlation skymap is the correlation score between two waveforms,
242 given some particular [time delay](#time_delays.make_time_delay_skymap), ie. phase shift.
244 
246 * `masks` can be used to mask the correlation maps, as shown in the bottom subplot in the
247 Figure above. The masks are defined by #supply_azimuthal_angle_masks.
249 \anchor scipy_corr_expl
251 * Consider two waveforms,
252 \anchor maxcorrachieved
253 \image html shift_by_472_samples.png
254 * Each row in the `correlation` column is an array of correlation scores.
255 * By shifting the waveforms we may be able to get them to align perfectly,
256 at which point maximum correlation is achieved.
257 * The correlation score tells us how "aligned" the two waveforms are after we phase shift
258 `A1_waveforms (volts)` against `A2_waveforms (volts)` by a certain amount of time.
259 * The waveforms are zero-centered and normalized such that the cross-correlation is
260 bounded between [-1,1]. Zero means the waveforms are not aligned at all.
261 * See [cross_correlation_and_time_delay.pdf](cross_correlation_and_time_delay.pdf) or,
262 for details, [scipy_correlate_behavior.pdf](scipy_correlate_behavior.pdf).
265 from scipy.signal
import correlate
267 correlation_maps: pl.DataFrame = (
271 pl.struct(
"A1_waveforms (volts)",
"A2_waveforms (volts)")
276 (wf1 - np.mean(wf1)) / np.std(wf1),
277 (wf2 - np.mean(wf2)) / np.std(wf2),
280 for wf1, wf2
in s.to_numpy()
283 .alias(
"correlation"),
287 pl.struct(pl.col(
"time delays [samples]"), pl.col(
"correlation")).map_batches(
289 [correlation_array[time_delay_matrix]
290 for time_delay_matrix, correlation_array
293 ).alias(
"correlation maps")
297 return correlation_maps
300def _get_true_direction(dataset: ROOT.pueo.Dataset) -> [float, float]:
302 @brief Returns the true signal direction in MC
304 @param[in] dataset The output of #initialise.load_pueoEvent_Dataset
306 * Note that as stored in the `.root` files,
307 the variable `RFdir_payload` is the direction the signal is travelling **to**.
308 * Therefore, to obtain the direction that the signal is coming **from**,
309 we need the opposite vector.
310 * Thus, \f$\phi_{\rm true} = (\phi_{\rm rfdir} + 180 ^\circ) \% 360^\circ\f$,
311 and \f$\theta_{\rm true} = 180^\circ - \theta_{\rm rfdir}\f$
314 truePhi = (dataset.truth().payloadPhi + 180) % 360
315 trueTheta = 180 - dataset.truth().payloadTheta
317 return truePhi, trueTheta
319def _get_true_pulser_direction_tmp(dataset: ROOT.pueo.Dataset, pulser_station:str, nav_file =
"../abxtwo.root") -> [float, float]:
321 @brief Temperary before payload gps data is attached to each event. Returns the true signal direction in payload coor.
323 @param[in] dataset The output of #initialise.load_pueoEvent_Dataset
324 @param[in] pulser_station name of the pulser station, ldb, taylor_dome, s200
325 @retval phi, theta in payload coor.
327 * Note that as stored in the `.root` files,
328 the variable `RFdir_payload` is the direction the signal is travelling **to**.
329 * Therefore, to obtain the direction that the signal is coming **from**,
330 we need the opposite vector.
331 * Thus, \f$\phi_{\rm true} = (\phi_{\rm rfdir} + 180 ^\circ) \% 360^\circ\f$,
332 and \f$\theta_{\rm true} = 180^\circ - \theta_{\rm rfdir}\f$
334 sys.path.append(
"/home/yuchiehku/pueo/pueoBuilder/components/pueoAnalysisTools")
335 import geographyUtils.locations
336 import geographyUtils.navigation
337 import geographyUtils.geometry
339 readout_time = dataset.header().readoutTime
341 locations_helper = geographyUtils.locations.LocationsHelper()
342 navigation_helper = geographyUtils.navigation.NavigationHelper(root_file=nav_file, root_source=
"ABX-Two")
343 geometry_helper = geographyUtils.geometry.GeometryHelper()
345 if len(pulser_station) > 0:
346 pulser_position = locations_helper.get_ground_pulser_position(pulser_station)
347 pueo_position = np.zeros(3)
348 pueo_position[0] = navigation_helper.get_longitude(readout_time, max_time_delta=1)
349 pueo_position[1] = navigation_helper.get_latitude(readout_time, max_time_delta=1)
350 pueo_position[2] = navigation_helper.get_altitude(readout_time, max_time_delta=1)
351 pueo_heading = navigation_helper.get_heading(readout_time, max_time_delta=1) + 45.3
352 expected_el_az = geometry_helper.get_direction_of_position(
357 expected_signal_direction = np.zeros(2)
358 expected_signal_direction[0] = expected_el_az[0]
359 expected_signal_direction[1] = 1*(-expected_el_az[1] + pueo_heading)
360 while expected_signal_direction[1] < 0:
361 expected_signal_direction[1] += 360
362 while expected_signal_direction[1] > 360:
363 expected_signal_direction[1] -= 360
365 expected_signal_direction =
None
367 return expected_signal_direction[1], -1*expected_signal_direction[0]+90
369def _get_true_pulser_direction(dataset: ROOT.pueo.Dataset, pulser_station:str) -> [float, float]:
371 @brief Returns the true signal direction from pulser and payload location
373 @param[in] dataset The output of #initialise.load_pueoEvent_Dataset
374 @param[in] pulser_station name of the pulser station, ldb, taylor_dome, s200
375 @retval phi, theta in payload coor.
377 * Does this function work on the data format? probably not now, but eventually it will
378 * Note that as stored in the `.root` files,
379 the variable `RFdir_payload` is the direction the signal is travelling **to**.
380 * Therefore, to obtain the direction that the signal is coming **from**,
381 we need the opposite vector.
382 * Thus, \f$\phi_{\rm true} = (\phi_{\rm rfdir} + 180 ^\circ) \% 360^\circ\f$,
383 and \f$\theta_{\rm true} = 180^\circ - \theta_{\rm rfdir}\f$
386 sys.path.append(str(Path(__file__).resolve().parent.parent))
387 import geographyUtils.locations
388 import geographyUtils.geometry
390 geometry_helper = geographyUtils.geometry.GeometryHelper()
391 locations_helper = geographyUtils.locations.LocationsHelper()
393 pulser_position = locations_helper.get_ground_pulser_position(pulser_station)
395 pueo_position = np.zeros(3)
398 pueo_position[0] = dataset.gps().longitude
399 pueo_position[1] = dataset.gps().latitude
400 pueo_position[2] = dataset.gps().altitude
402 pueo_heading = dataset.gps().heading
409 expected_el_az = geometry_helper.get_direction_of_position(
414 expected_signal_direction = np.zeros(2)
417 expected_signal_direction[0] = 180 - expected_el_az[0]
420 expected_signal_direction[1] = (-expected_el_az[1] + 180 + pueo_heading)
422 while expected_signal_direction[1] < 0:
423 expected_signal_direction[1] += 360
424 while expected_signal_direction[1] > 360:
425 expected_signal_direction[1] -= 360
427 truePhi = expected_signal_direction[1]
428 trueTheta = expected_signal_direction[0]
430 return truePhi, trueTheta
432def plot_correlation_map(correlation_frame: pl.DataFrame, plot_name: str,
433 true_phi=
None, true_theta=
None) ->
None:
435 @brief Plots the reult of #cross_correlate.
437 @param[in] correlation_frame The output of #cross_correlate
438 @param[in] plot_name Remember to specify file type
439 @param[in] true_phi (optional) from #_get_true_direction
440 @param[in] true_theta (optional) from #_get_true_direction
442 * Required columns in `correlation_frame`:
443 -# `correlation maps`
447 * Using only one antenna pair, one can find a band of peak correlation scores,
448 see the [plot in the file description](@ref example_one_pair_corr_map).
450 * If we then sum over all antenna pairs, we would be able to identify a single peak:
452 
455 import matplotlib.pyplot
as plt
456 import numpy.ma
as ma
457 plt.rcParams.update({
'font.size': 15})
459 phi, theta = _make_grid(sparse=
True)
460 phi = np.degrees(phi)
461 theta = np.degrees(theta)
463 fig, axes = plt.subplots(2, 1, figsize=(20, 10), sharex=
True)
464 for i, df
in enumerate(correlation_frame.partition_by(
"Pol")):
465 corr_map = df[
"correlation maps"].to_numpy()
468 masks = np.repeat(df[
"masks"].to_numpy(), repeats=np.shape(corr_map)[1], axis=1)
469 masks_sum = np.sum(~masks, axis=0)
471 corr_map_masked = ma.array(corr_map, mask=masks)
474 total_masked = np.sum(corr_map_masked, axis=0) / masks_sum
476 axes[i].set_ylabel(
r"$\theta$")
477 axes[i].set_xlabel(
r"$\phi$")
478 axes[i].set_title(f
"{df['Pol'][0]}Pol correlation maps")
479 fig.colorbar(axes[i].pcolormesh(phi, theta, total_masked))
481 if true_phi
is not None and true_theta
is not None:
482 axes[i].scatter(true_phi, true_theta, marker=
'x', color=
"red", label=
'True Direction')
485 fig.savefig(plot_name)
489def __plot_example_correlation_map_for_one_antenna_pair(input_frame: pl.DataFrame):
490 import matplotlib.pyplot
as plt
491 import numpy.ma
as ma
492 plt.rcParams.update({
'font.size': 15})
494 pl.Config().set_tbl_cols(input_frame.width)
495 pl.Config().set_fmt_table_cell_list_len(0)
499 mask = input_frame[
"masks"].to_numpy().squeeze()
500 corr_map = input_frame[
"correlation maps"].to_numpy().squeeze()
503 mask = np.vstack([mask] * corr_map.shape[0])
504 corr_map_masked = ma.array(corr_map, mask=mask)
507 phi, theta = _make_grid(sparse=
True)
509 fig, axes = plt.subplots(2, 1, sharex=
True, figsize=(20, 10))
511 f
"Antennas {input_frame['A1_AntNum'][0]} and {input_frame['A2_AntNum'][0]} Correlation Map"
514 ax.set_ylabel(
r"$\theta$")
515 ax.set_yticks([np.pi / 2, np.pi], labels=[
r"$\pi / 2$",
r"$\pi$"])
517 axes[1].set_xlabel(
r"$\phi$")
518 axes[1].set_xticks([0, np.pi, 2 * np.pi], labels=[
"0",
r"$\pi$",
r"$2\pi$"])
519 image = axes[0].pcolormesh(phi, theta, corr_map, vmin=corr_map.min(), vmax=corr_map.max())
520 image = axes[1].pcolormesh(phi, theta, corr_map_masked, vmin=corr_map.min(), vmax=corr_map.max())
521 fig.colorbar(image, ax=axes.ravel().tolist(), label=
"Correlation Score")
522 plt.savefig(
"img/example_correlation_map_one_pair.png")
525if __name__ ==
"__main__":
527 from antenna_attributes
import read_antenna_geometry, get_nominal_phase_center
528 from antenna_pairs
import generate_MI_antenna_pairs
529 from time_delays
import make_time_delay_skymap
530 from waveform_plots
import load_waveforms, upsample_waveforms
534 _jun25: Path = os.environ.get(
"PUEO_UTIL_INSTALL_DIR") / Path(
"share/pueo/geometry/jun25/qrh.dat")
535 _face_centers: pl.DataFrame = read_antenna_geometry(qrh_dot_dat=_jun25)
537 _phase_centers: pl.DataFrame = (
538 get_nominal_phase_center(face_centers=_face_centers)
539 .select(
"AntNum",
"PhiSector",
"AntIdx",
"X[m]",
"Y[m]",
"Z[m]")
544 _antenna_pairs: pl.DataFrame = generate_MI_antenna_pairs(antennas=_phase_centers)
545 _time_delays: pl.DataFrame = (
546 make_time_delay_skymap(antenna_pairs=_antenna_pairs)
547 .select(pl.col(
r"^A[12]_AntNum$",
r"^A[12]_PhiSector$",
"time delays [sec]"))
552 masked: pl.DataFrame = (
553 supply_azimuthal_angle_masks(skymaps=_time_delays)
554 .select(pl.col(
r"^A[12]_AntNum$",
"time delays [sec]",
"masks"))
559 _run_zero_data: ROOT.pueo.Dataset = load_pueoEvent_Dataset(pueo_mc_data=Path(
"/tmp"), run_number=0)
560 _run_zero_data.last()
562 _wf: pl.DataFrame = (
563 load_waveforms(dataset=_run_zero_data)
564 .select(
"AntNum",
"Pol",
"waveforms (volts)",
"step size (nanoseconds)")
566 up: pl.DataFrame = upsample_waveforms(waveforms=_wf, upsample_factor=3)
570 big_frame = combine_time_delay_maps_and_waveforms(masks_and_skymaps=masked, waveforms=up)
572 correlation_frame: pl.DataFrame = cross_correlate(big_frame)
576 tp, tt = _get_true_direction(dataset=_run_zero_data)
577 plot_correlation_map(
578 correlation_frame=correlation_frame, true_phi=tp, true_theta=tt,
579 plot_name=
'img/example_correlation_map_two_pols.png'